In this post we will journey on an expedition to explore a forestry dataset using Data Science tools available in Python. Python is an Object-Oriented programming language commonly used in the Geospatial and Data Science fields. As an example scenario, we will work with two datasets that resemble common Stand and Stocking attribute data exported from a GIS. We will use Python in a Jupyter Notebook environment, and explore the dataset using an Exploratory Data Analysis (EDA) approach.
The Engineering Statistics Handbook says that the primary goal of Exploratory Data Analysis is to “maximize the analyst’s insight into a data set and into the underlying structure of a data set…”. An EDA approach examines the dataset first, allowing the analyst to determine which model would best represent the data. Contrast this with the classical statistical approach which would first impose a hypothesis, or model, prior to seeing the data. This is about as deep into the theory as we will get, but just know that when analyzing a dataset, EDA is often the first phase of the analytic process.
EDA employs both graphical and non-graphical methods for exploring datasets. In the example that follows we will look at both. We will see that Python, and a commonly associated data science library called Pandas, provide efficient methods for working with datasets. Much of this analysis could be done using a Spreadsheet program such as Microsoft Excel. But, I prefer working with Python and Pandas in a Jupyter Notebook environment, because I find the process more efficient, easily reproducible, and easier to spot issues within the dataset.
Example EDA Process
For the example dataset we will look at data collected from a managed forest Geographic Information System (GIS) in two formats. The first dataset is an ESRI feature class in SHP format containing forest stand attribute data. This dataset will be opened using GeoPandas – a geographic variant of Pandas. The second dataset contains Stand Inventory records in CSV (comma-separated-values) format, which we will open using Pandas.
It should be noted that both of these datasets are “tidy”, where each variable is a column, each observation is a row, and the combination of rows and columns forms a table. Also, the data is mostly clean, so we will begin by combining the datasets into one.
Restructuring the Data
Before we begin analysis of these datasets, we need to restructure the data. Currently, the data is in two separate datasets. For our analysis, we need to combine these into one.
First we will import the required packages in our Jupyter Notebook environment. If you are not familiar with Jupyter Notebooks you can find more info about them here.
# import the required packages
import pandas as pd
import geopandas as gpd
import matplotlib.pyplot as plt
import seaborn as sns; sns.set()
Next, we read the datasets into stored variables using the GeoPandas and Pandas libraries. This will allow us to do additional processing on the datasets, without having to import them each time. The filename of the stand data is ‘planted_stands.shp’ , and it is located in the current notebook path. The filename of the inventory data is ‘stocking.csv’ and it is also in the current notebook path. The object we are using to store the records is called a DataFrame. A DataFrame is a 2-dimensional data structure which is similar to an Excel Spreadsheet with rows and columns. The data is represented as features (columns) and observations (rows).
# import the SHP into GeoPandas DataFrame
planted_stands = gpd.read_file('planted_stands.shp')
# import the CSV into Pandas DataFrame
stocking_data = pd.read_csv('stocking.csv')
# show the first 5 records of each dataset
planted_stands.head()
stocking_data.head()
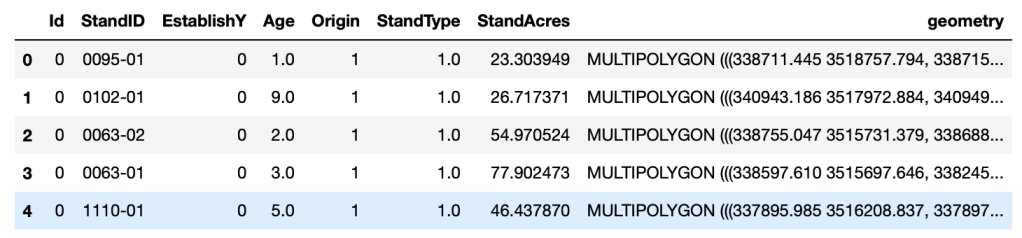
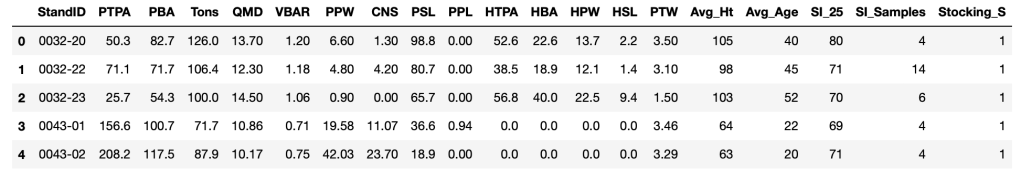
Next, we will convert the GeoPandas DataFrame into a regular Pandas DataFrame. We do this because we do not need the Geographic attributes of the dataset (i.e.; ‘geometry’) for this analysis, and we prefer the simplicity of working with a single datatype.
# drop the geometry column and convert this to a Pandas DataFrame
planted_data = pd.DataFrame(planted_stands.drop(column='geometry'))
The resulting DataFrame is shown below. Notice that the geometry column has now been removed.
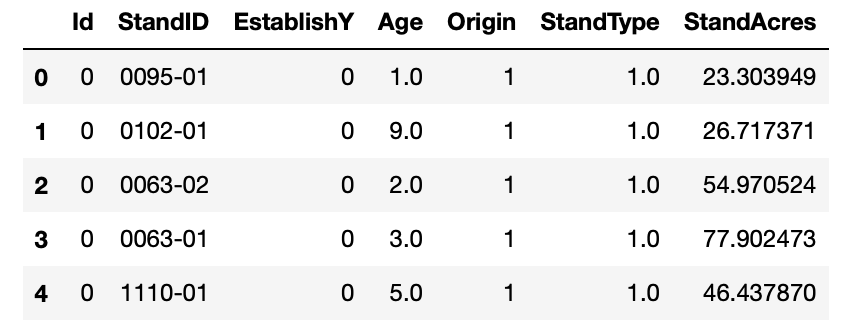
At this point, we can merge the two DataFrames into one, which will make it easier for the analysis phase. The merge() function in Pandas is similar to a JOIN in SQL database language, with the default being an “INNER JOIN” on a specified column.
# merge the DataFrames into one using the default "inner"
full_data = pd.merge(planted_data, stocking_data, on='StandID')
The DataFrames were merged on the ‘StandID’ column. This feature was selected because it represents a unique stand observation in each dataset. The goal here is to join matching stand records in both DataFrames. In other words, if stand “0095-01” exists in both DataFrames, then these records will be combined. Here is the result of the merge operation on the DataFrames.
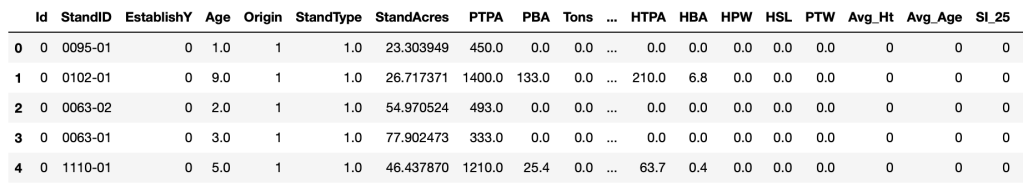
Analysis
Now that the datasets are combined, we can begin the analysis. The first step here will be to calculate some summary statistics on the dataset. We will use the Pandas describe() method on the DataFrame object for this step.
# get simple stats on the data
full_data.describe()
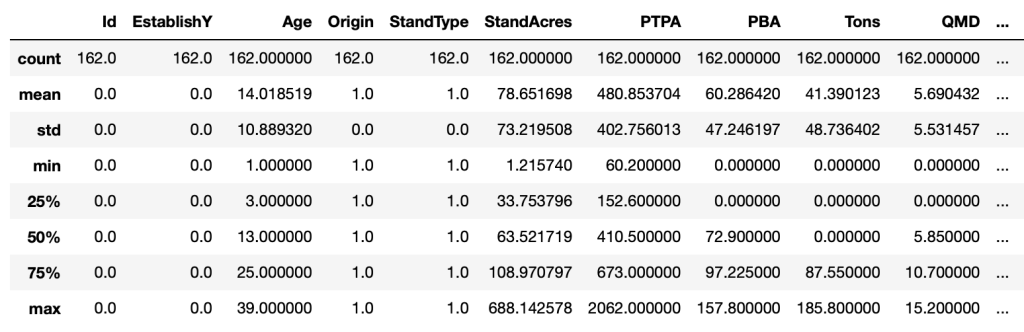
The describe method provides descriptive statistics for each of the features in our DataFrame. We can see that the average stand age is 14 years old, ranging from 1 (min) to 39 (max), and the average stand acreage is 78.6, ranging from 1.2 (min) to 688.1 (max). The result of the count operation shows that there are 162 values in each of the features, which indicates that there are no “NaN” values in this dataset. NaN is a special value that means “not-a-number”, or in other words missing data.
Another exploratory operation provided by Pandas is the info() method, which will return additional metadata about the DataFrame.
# view metadata about the DataFrame
full_data.info()
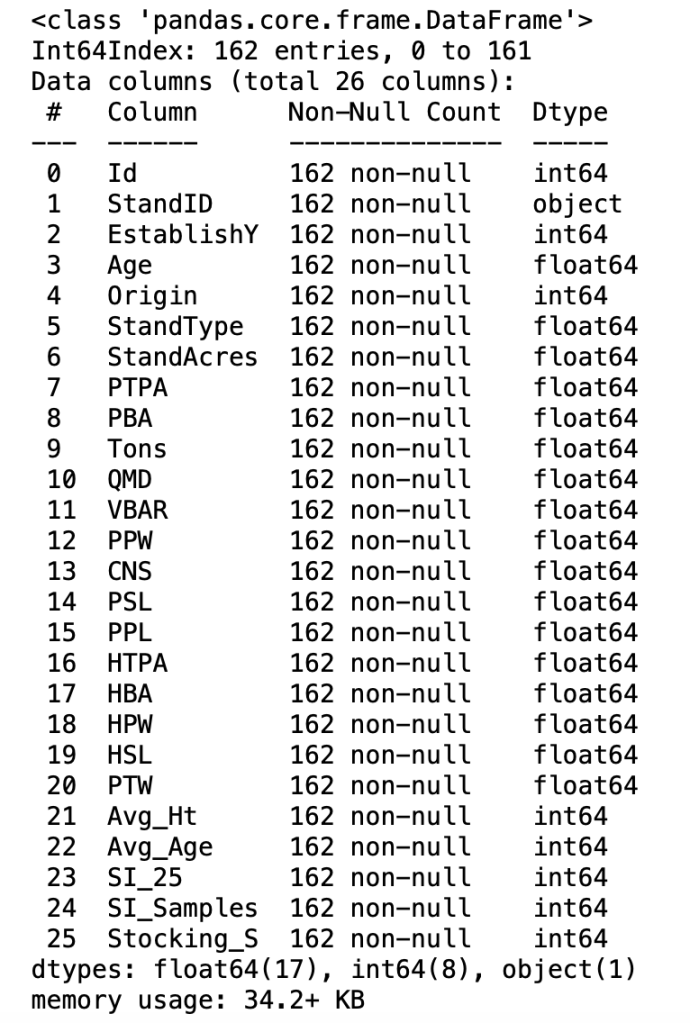
Pandas will return some basic metadata about the DataFrame including the feature (Column) names, the datatype, and the amount of computer memory (i.e.; RAM) occupied by the DataFrame. The info method will also show the number of “non-null” data values for each column (e.g.; 162), which is an alternative to the count operation of the describe method discussed above.
Another exploratory step involves calculating the correlations between features in the dataset. Because there are a lot of features in this dataset, we will filter out the ones that are of most interest to us using DataFrame indexing. Indexing is a term that basically means returning a partial view of the DataFrame. In this case we will only return a subset of features and perform correlation calculations on each of them using the corr() method. The correlation values will be shown as annotations in a heat map. Graphical analysis is a very productive form of EDA.
# select specific features to assess correlations
cols = ['PTPA', 'PBA', 'QMD', 'Tons', 'Avg_Age', 'Avg_Ht']
part_data = full_data[cols]
fig, ax = plt.subplots(figsize=(8, 8))
ax = sns.heatmap(part_data.corr(), cmap='YlGnBu', annot=True)
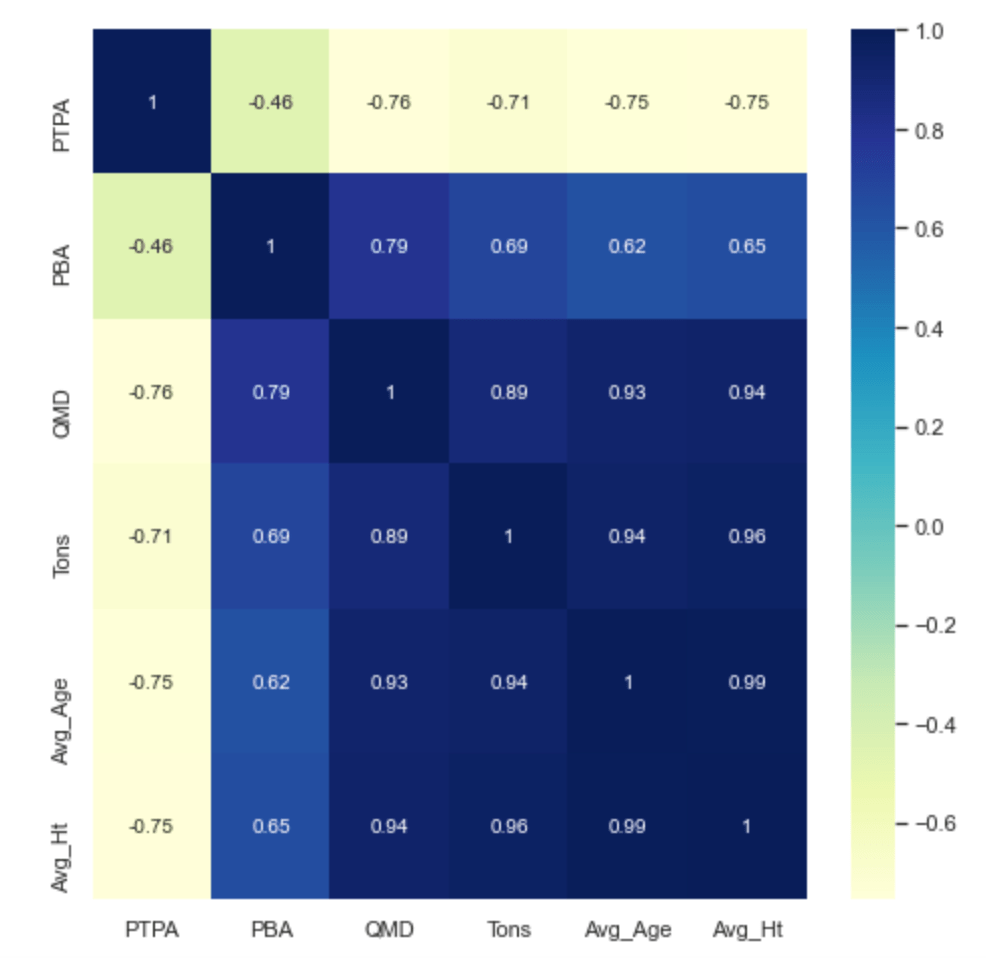
The correlation calculations shown above reveal that QMD (Quadratic Mean Diameter) has a strong positive correlation with Tons per acre, and that Trees per acre have a strong negative correlation with QMD. In other words, as diameter increases and trees per acre decreases, tons per acre tend to also increase. This is a commonly observed relationship in Forestry and hasn’t really revealed any new information, nevertheless it is a good exercise to see how the features are correlated. Remember “correlation does not imply causation“, rather it’s an indicator of a relationship, and how strong or weak that relationship is.
Now, we will plot a few of these relationships to see if there is something we can learn from the data.
# plot Pine TPA by Pine BA
fig, ax = plt.subplots(figsize=(6, 6))
plt.scatter(full_data['PTPA'], full_data['PBA'], s=50)
plt.xlabel('Pine Trees per Acre')
plt.ylabel('Pine Basal Area per Acre')
plt.show()
We will plot the Basal Area per acre over Trees per acre (also known as a Stocking Chart) using the scatter plot method provided by the Matplotlib library. Matplotlib is a graphical library for producing visualizations in Python. Below is the resulting Stocking Chart.
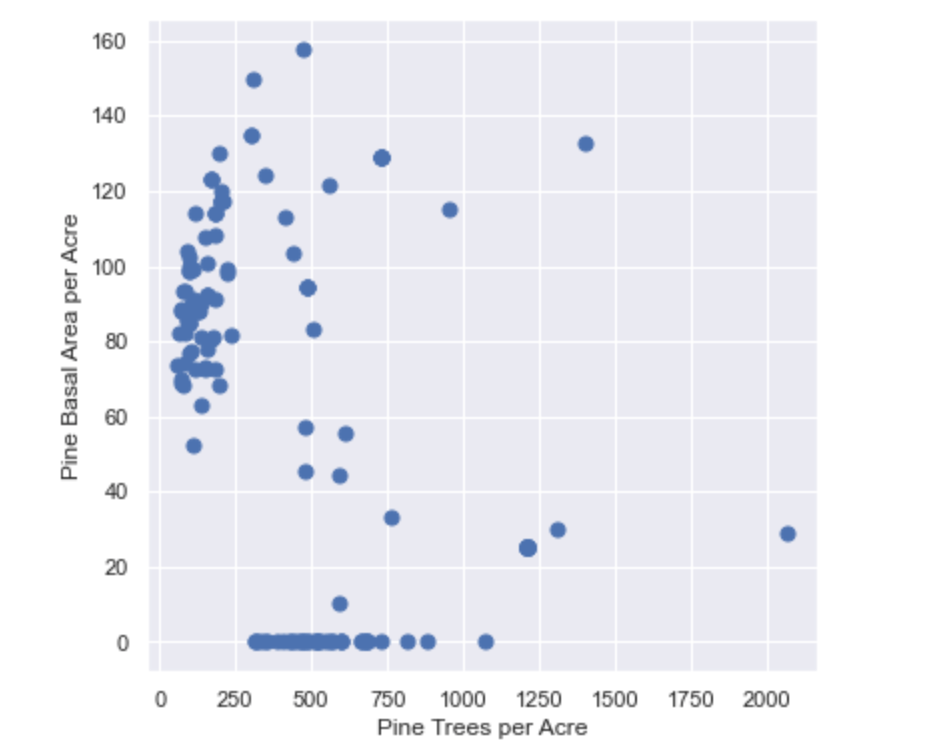
One thing to notice about this chart is that there are several stands with no reported Basal Area. This is because these stands are juvenile (< 10 years old) and have not been assigned a Basal Area value yet. Therefore, it’s probably best to remove them from the DataFrame and recreate the chart.
# create a new dataset which includes stands greater than age 9
stands_gt10 = full_data[full_data['Age'] > 9]
The above code created a new DataFrame called ‘stands_gt10’ by filtering the previous DataFrame using the condition, age is greater than 9 years. We will use this DataFrame in the analysis from here out. Let’s recreate the previous plot of Basal Area per Acre over Trees per Acre now that the juvenile stands have been removed.
# plot a scatterplot of Pine TPA by Pine BA
fig, ax = plt.subplots(figsize=(6, 6))
plt.scatter(stands_gt10['PTPA'], stands_gt10['PBA'], s=50)
plt.xlabel('Pine Trees per Acre')
plt.ylabel('Pine Basal Area per Acre')
plt.show()
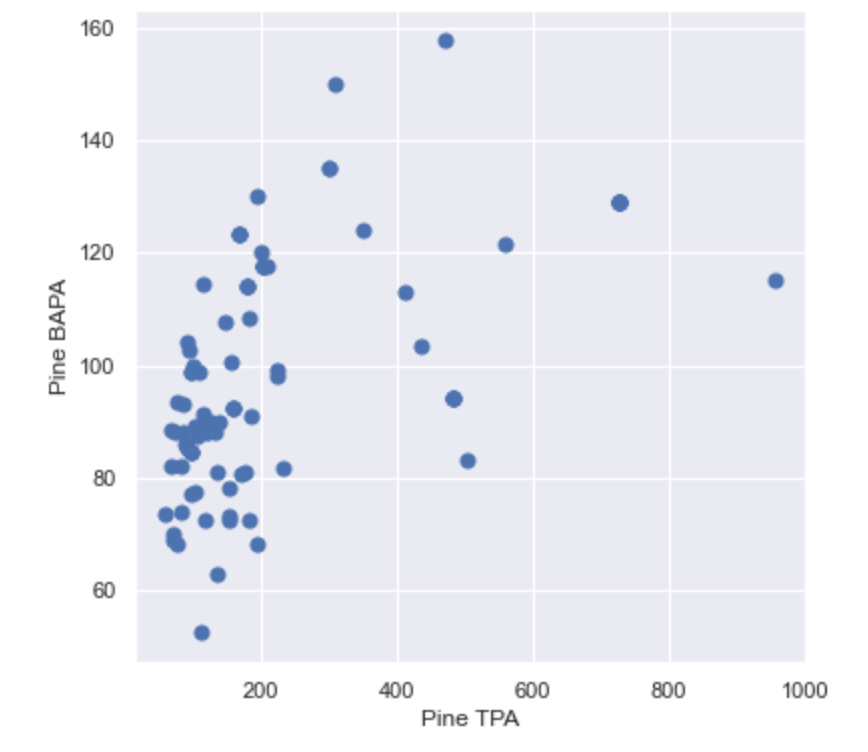
This version of the stocking chart shows that most of the merchantable plantations are clustered below 200 Trees per Acre and 120 Basal Area per Acre. It also shows outliers, which indicate there may a few overstocked or understocked stands, but overall these are “well-stocked”. In a similar plot, we will observe QMD and Trees per Acre, which is the relationship that Stand Density Index (SDI) is based upon.
To create this plot we need to do a little more work with the data by creating X and Y values for the SDI baselines. We will superimpose these onto the plot.
# plot QMD over TPA
import math
fig, ax = plt.subplots(figsize=(8,6))
# x, y values for 50% max SDI
y_vals50 = range(15,2,-1)
x_vals50 = [10 ** (math.log(225, 10) + 1.605 - (1.605 * math.log(y, 10))) for y in y_vals]
# x, y values for 35% max SDI
y_vals35 = range(15,2,-1)
x_vals35 = [10 ** (math.log(158, 10) + 1.605 - (1.605 * math.log(y, 10))) for y in y_vals]
plt.scatter(stands_gt10['PTPA'], stands_gt10['QMD'], alpha=0.8, s=50, c='red')
sdi50, = plt.plot(x_vals50, y_vals50, '--', c='b', linewidth=3, label='50% Max SDI')
sdi35, = plt.plot(x_vals35, y_vals35, '--', c='g', linewidth=3, label='35% Max SDI')
plt.xlabel('Pine TPA')
plt.ylabel('QMD')
plt.legend(handles=[sdi50, sdi35])
plt.show()
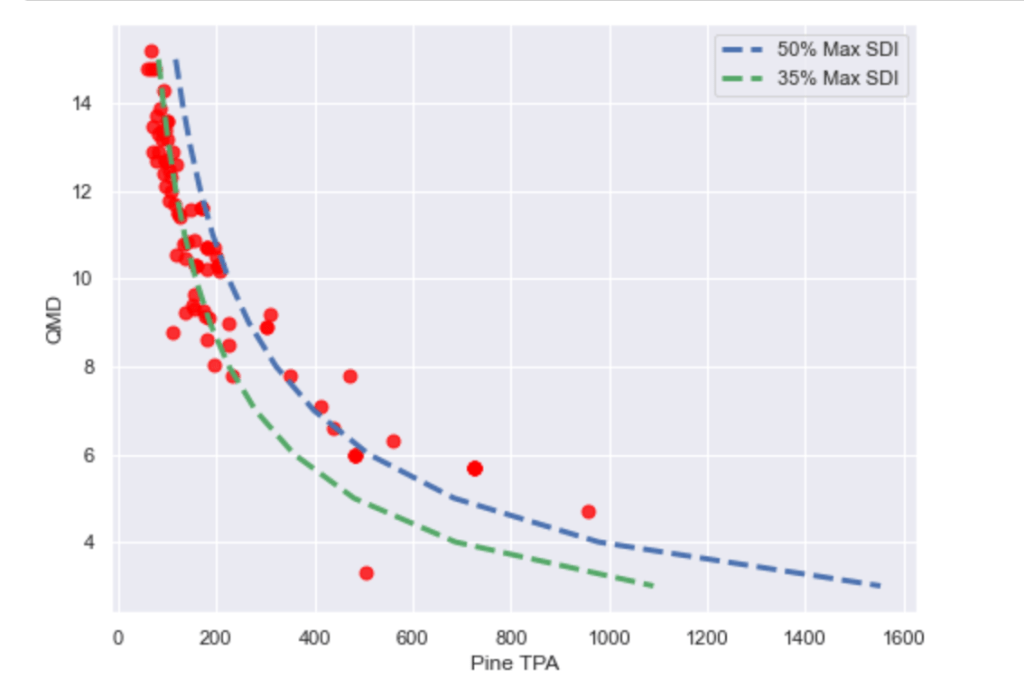
For this chart we developed SDI curves for 35% and 50% max SDI values for Loblolly Pine. Trees per Acre were calculated for each combination of SDI and QMD in the dataset. As this chart shows, most of the stands are below the 50% threshold, which indicates the threshold where self-thinning occurs. However, there are a few stands (~ eight) above the 50% threshold, which suggests that these stands need to be thinned as soon as possible.
This demonstrates the graphical approach, but we could have calculated the SDI for each stand and appended the values to a new column in the DataFrame. The code to accomplish this is shown below.
# calculate SDI and add a column to the data
import math
def sdi(tpa, bapa, qmd):
'''returns max sdi as a float'''
sdi = (qmd / 10)**1.605
return (sdi * tpa) / 450
stands_gt10['sdi'] = stands_gt10.apply(lambda row: sdi(row['PTPA'], row['PBA'], row['QMD']), axis=1)
stands_gt10
Here we are using a custom function called sdi with inputs of Trees per Acre, Basal Area per Acre and tree Diameter to calculate the SDI value. We apply this function to each row in the DataFrame and assign the resulting value to a new column in the DataFrame called stands_gt10[‘sdi’]. The following partial view of the DataFrame shows the “sdi” column appended to the end of the previous DataFrame.
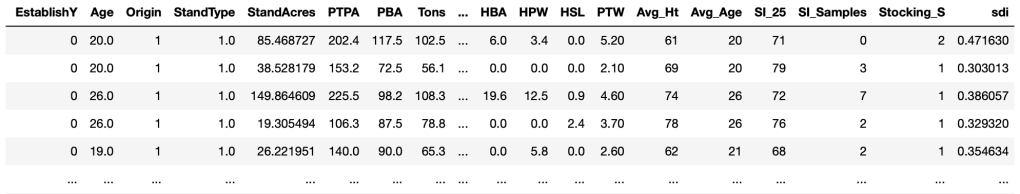
Data Frame showing sdi appended to the end.
Summary
This article introduced Exploratory Data Analysis by providing an example workflow on common Forestry datasets. We used Pandas for working with structured data as DataFrames, and Matplotlib for providing graphical presentations of the data. We could have done a much deeper dive into the dataset using linear and non-linear Regression and Classification techniques. Maybe this will be the subject of another article. Overall, I believe we saw a good example of combining data from multiple sources, and analyzing the data to see what simple patterns or trends exist.
Categories: Data Analytics
Excellent walk through. Please consider teaching, you have a gift!
LikeLike
Dude!!! You are so way over my head
Dallas Geis
580-351-4459
http://www.genesisindustries.net
LikeLike